Estimates the number of iterations needed for the null model randomization in
ps_rand() to reach its stationary distribution, given a dataset and algorithm.
This is a convenience wrapper around nullcat::suggest_n_iter() that extracts the
appropriate community matrix from a phylospatial object. Use this before running
ps_rand() with fun = "nullcat" or fun = "quantize" to choose an appropriate
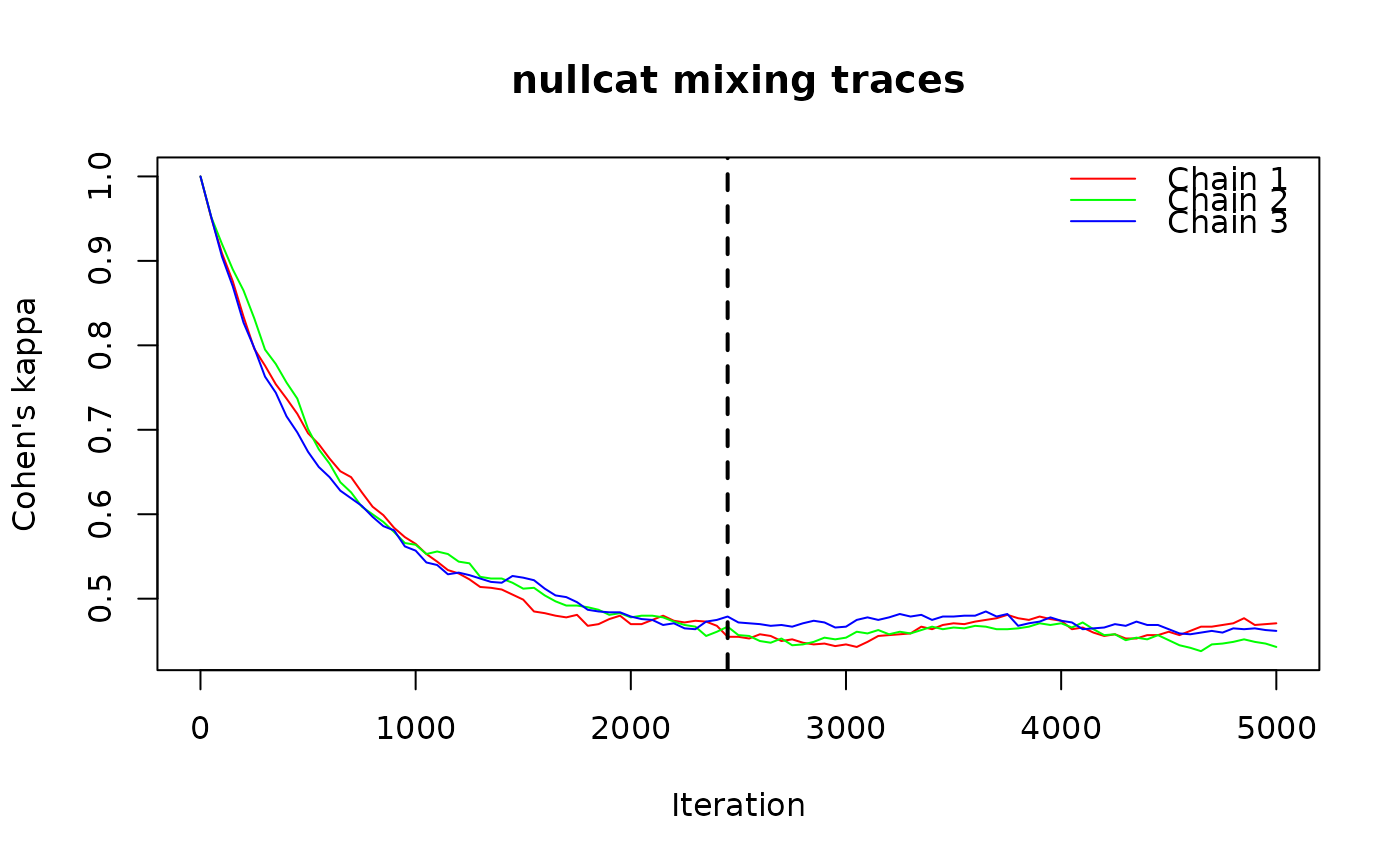
value for n_iter. The function runs multiple independent chains of the randomization
algorithm, records a mixing diagnostic at each step, and identifies the point at which
chains stabilize.
Usage
ps_suggest_n_iter(ps, fun = c("nullcat", "quantize"), plot = TRUE, ...)Arguments
- ps
A
phylospatialobject.- fun
Character:
"nullcat"or"quantize", matching the intendedfunargument tops_rand().- plot
Logical: if
TRUE, plot the mixing traces with the suggested burn-in marked.- ...
Additional arguments passed to
nullcat::suggest_n_iter()and through tonullcat::trace_cat(), such asmethod,n_iter,n_chains,n_strata,fixed, etc.
Value
An integer giving the suggested minimum number of iterations, with additional
diagnostic information as attributes. See nullcat::suggest_n_iter() for details.
Examples
# \donttest{
if (requireNamespace("nullcat", quietly = TRUE)) {
set.seed(123)
# binary data with nullcat
ps_bin <- ps_simulate(data_type = "binary")
ps_suggest_n_iter(ps_bin, fun = "nullcat", method = "curvecat",
n_iter = 5000, n_chains = 3, plot = TRUE)
# quantitative data with quantize
ps <- ps_simulate(data_type = "prob")
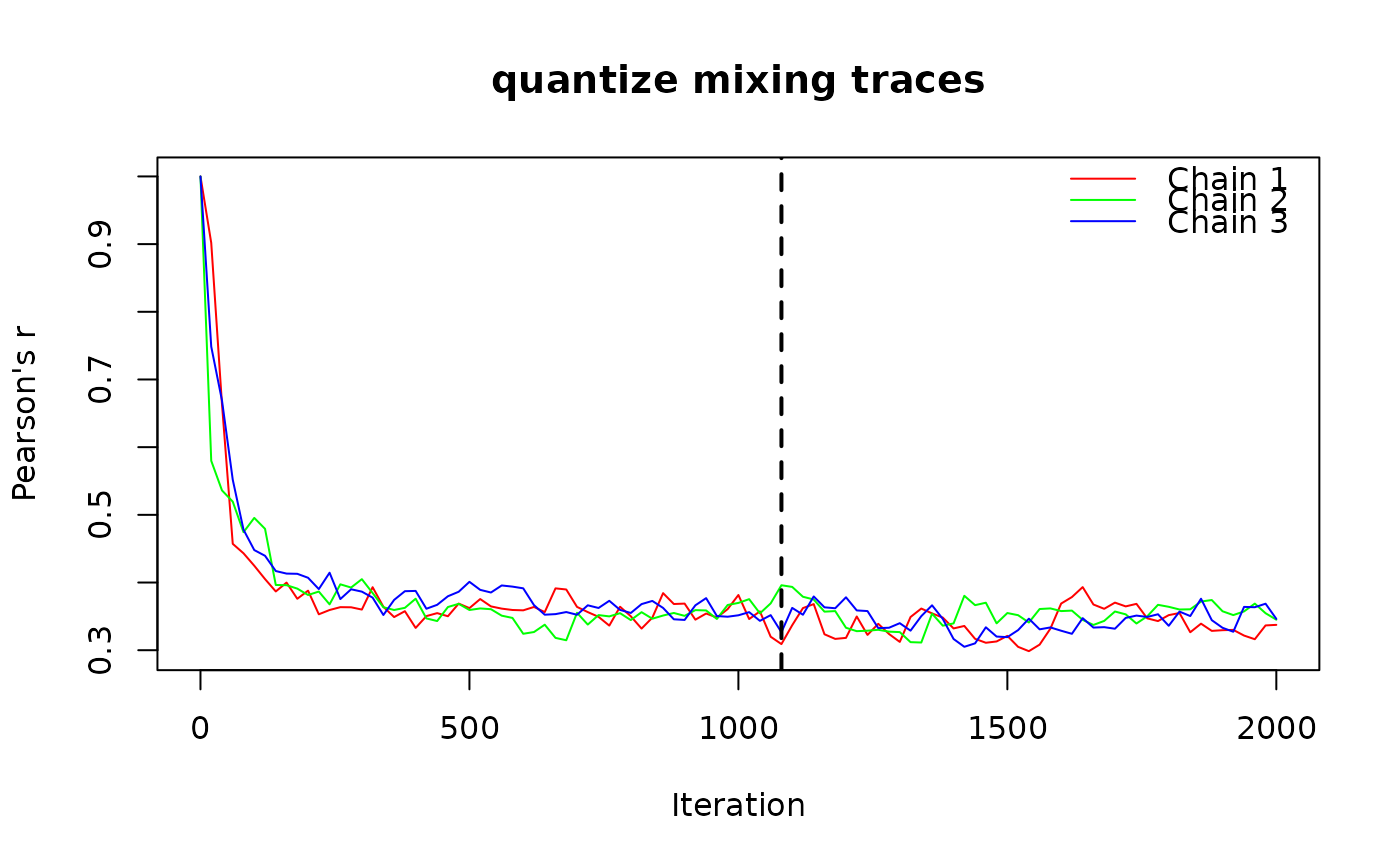
ps_suggest_n_iter(ps, fun = "quantize", method = "curvecat",
n_strata = 4, fixed = "cell",
n_iter = 2000, n_chains = 3, plot = TRUE)
}
 #> suggested_n_iter object
#> -----------------------
#> Converged: TRUE
#> Suggested n iterations: 1080
# }
#> suggested_n_iter object
#> -----------------------
#> Converged: TRUE
#> Suggested n iterations: 1080
# }
