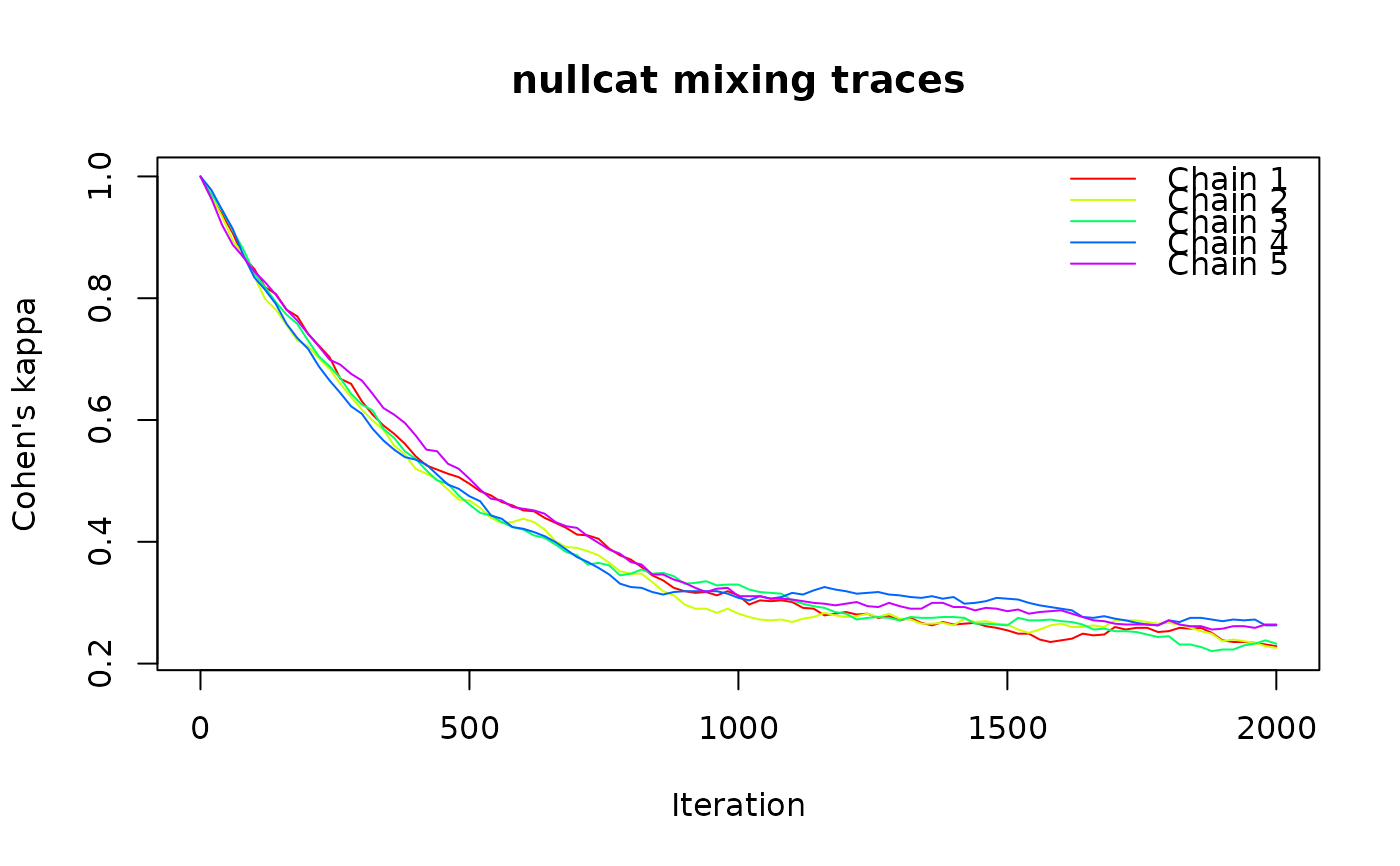
Runs multiple independent chains of the null model randomization algorithm and
records a mixing diagnostic at each step, producing trace plots to assess convergence.
This is a convenience wrapper around nullcat::trace_cat() that extracts the
appropriate community matrix from a phylospatial object.
Usage
ps_trace(ps, fun = c("nullcat", "quantize"), plot = TRUE, ...)Arguments
- ps
A
phylospatialobject.- fun
Character:
"nullcat"or"quantize", matching the intendedfunargument tops_rand().- plot
Logical: if
TRUE, plot the traces.- ...
Additional arguments passed to
nullcat::trace_cat(), such asmethod,n_iter,thin,n_chains,n_strata,fixed,stat, etc.
Value
An object of class "cat_trace". See nullcat::trace_cat() for details.
Examples
# \donttest{
if (requireNamespace("nullcat", quietly = TRUE)) {
ps_bin <- ps_simulate(data_type = "binary")
tr <- ps_trace(ps_bin, fun = "nullcat", method = "curvecat",
n_iter = 2000, n_chains = 5, plot = TRUE)
ps <- ps_simulate(data_type = "prob")
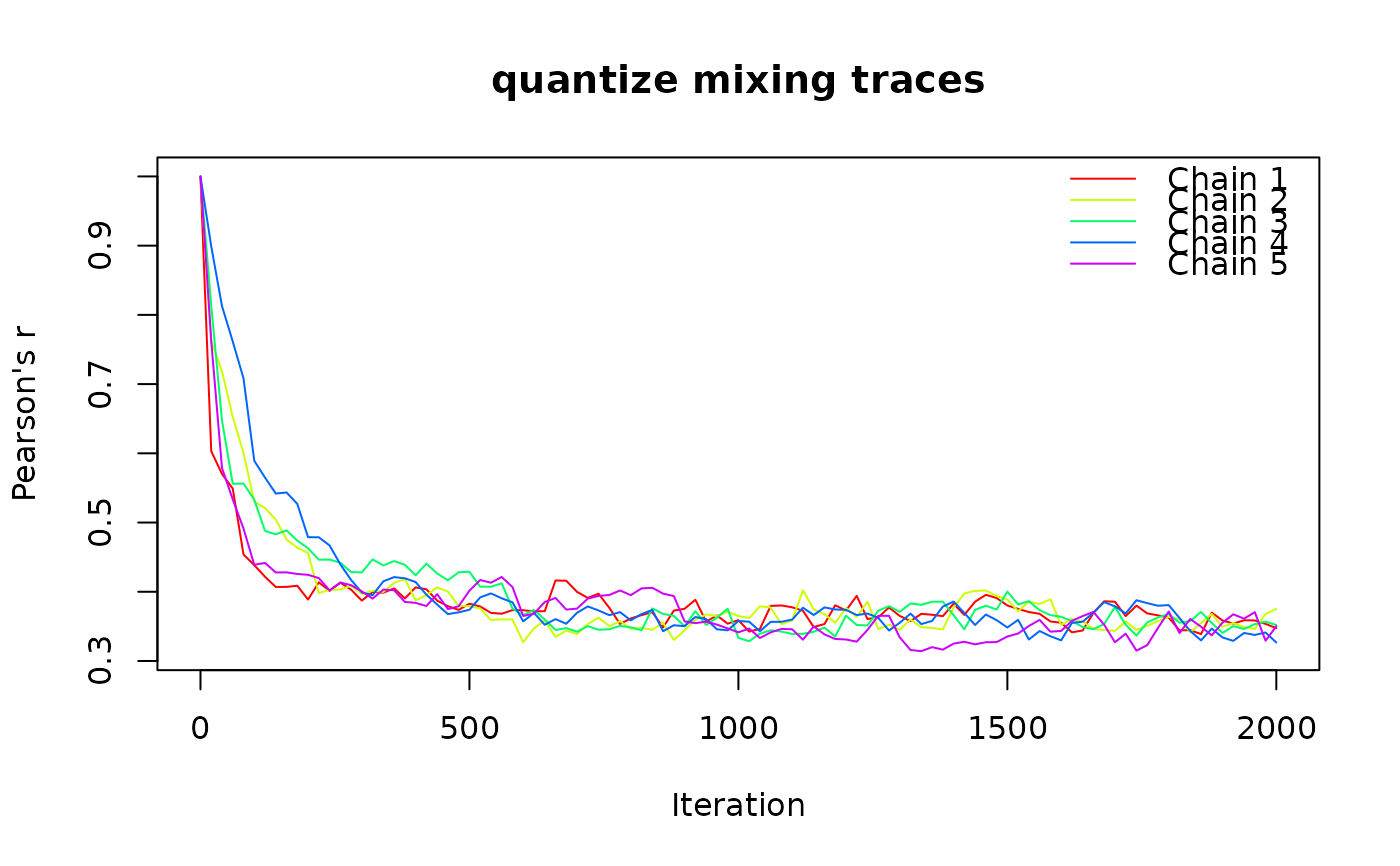
tr <- ps_trace(ps, fun = "quantize", method = "curvecat",
n_strata = 4, fixed = "cell",
n_iter = 2000, n_chains = 5, plot = TRUE)
}
 # }
# }
