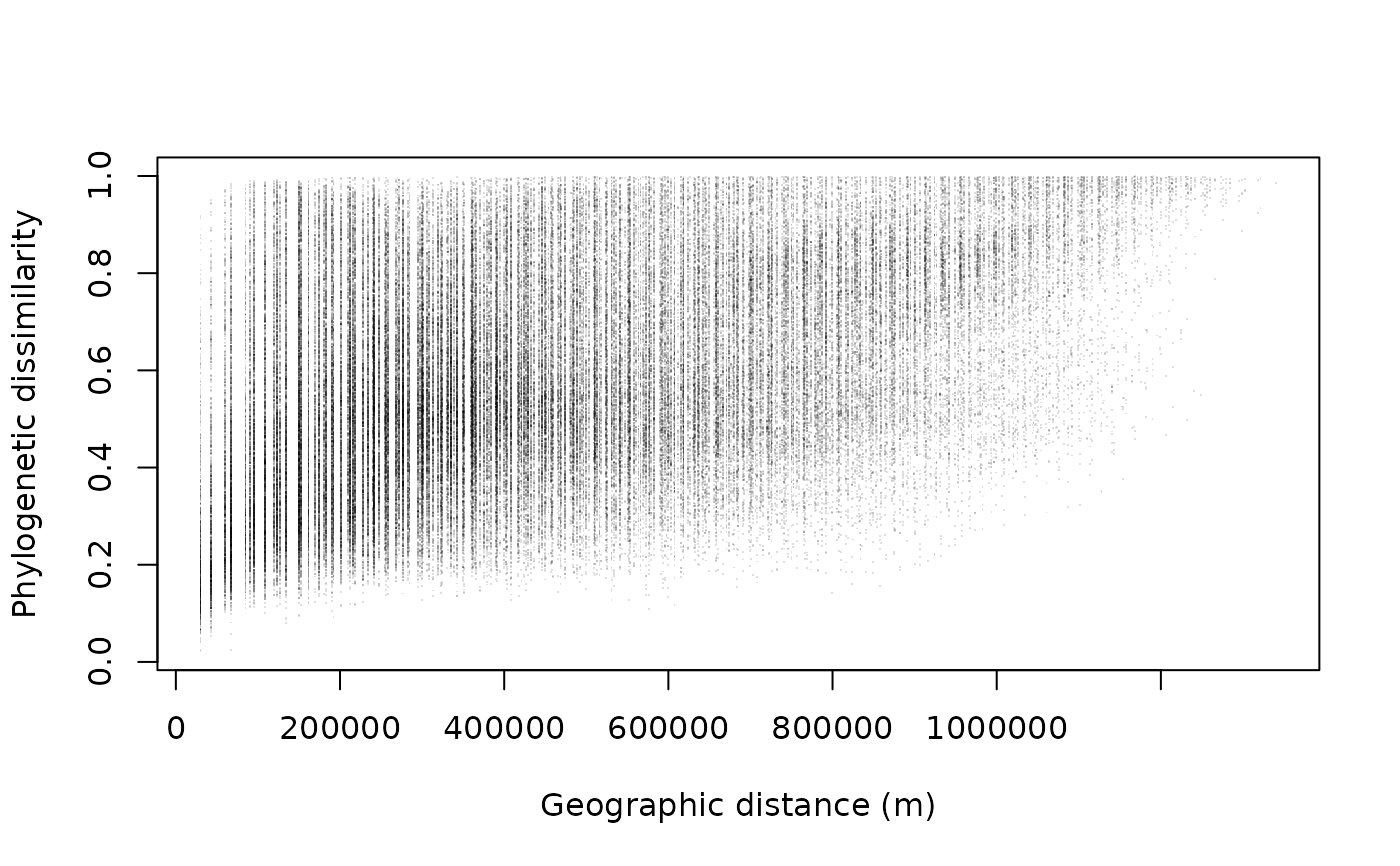
Calculate pairwise geographic distances between occupied sites in a phylospatial object.
The result is a dist object with the same dimensions and site ordering as ps_dissim(),
making it straightforward to compare phylogenetic dissimilarity with geographic distance
(e.g., for distance-decay analyses or Mantel tests).
Value
A pairwise geographic distance matrix of class dist, with one entry per pair
of occupied sites. Units are meters when a CRS is defined, or raw coordinate units otherwise.
Details
For raster data, distances are computed between cell centroids. For polygon or line sf data,
distances are computed between geometry centroids. Great-circle distances are used automatically
when the data has a geographic (lon/lat) CRS; Euclidean distances are used for projected data
or data without a CRS. Units are meters when a CRS is defined, or unitless coordinate
distances otherwise.